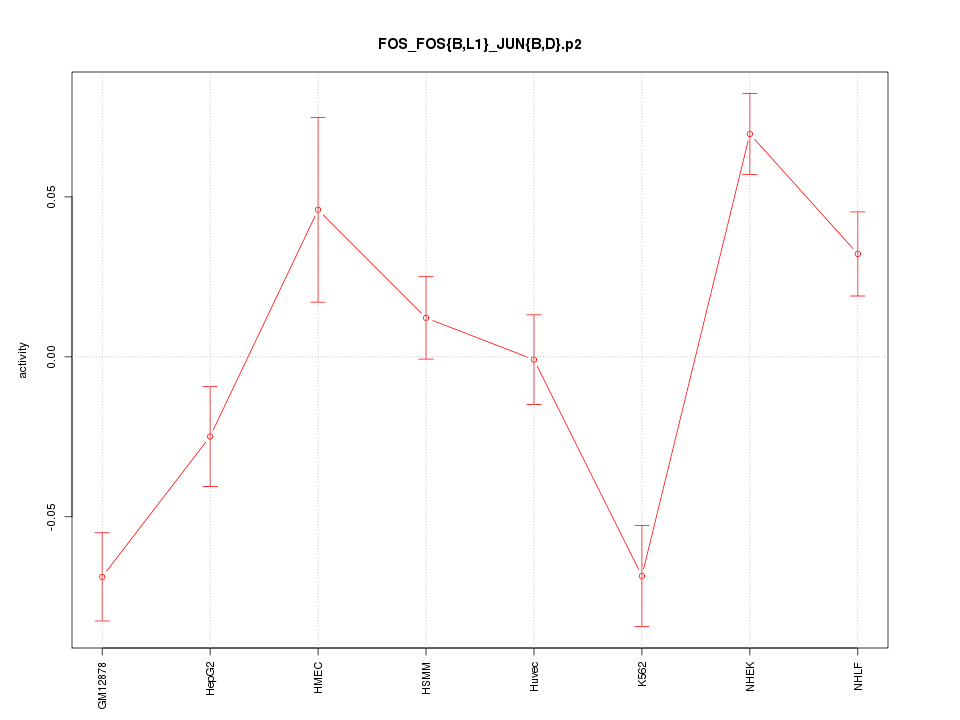
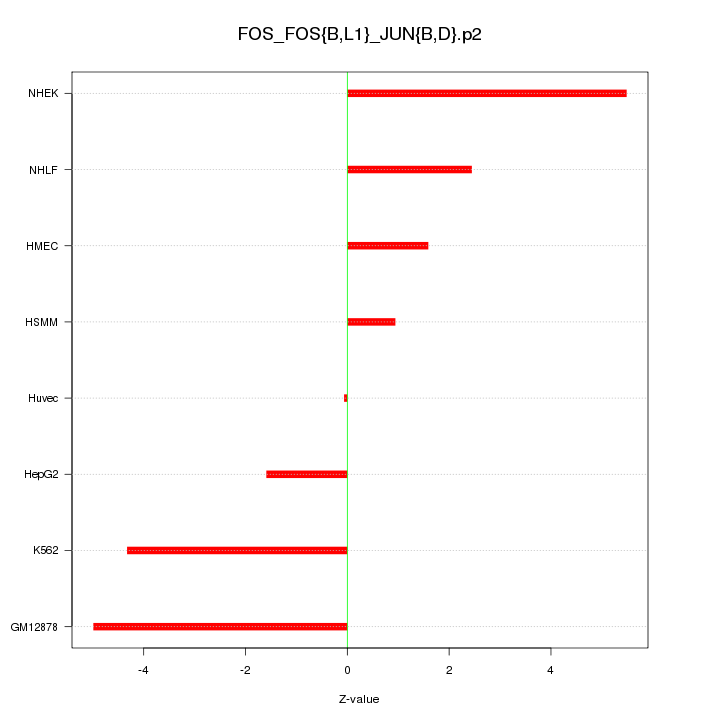
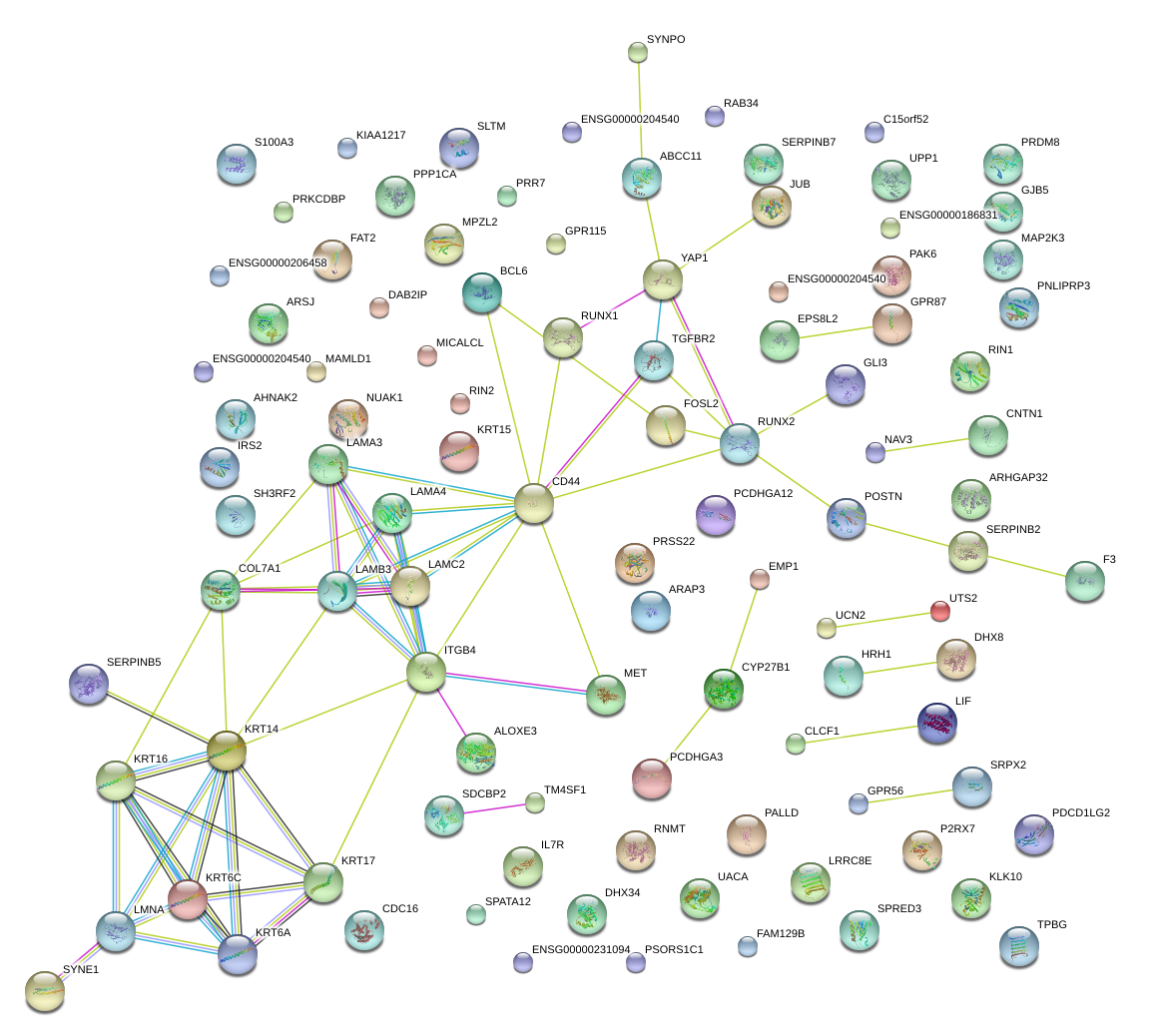
|
chr12_+_13240868
|
9.206
|
NM_001423
|
EMP1
|
epithelial membrane protein 1
|
|
chr12_+_13240987
|
8.057
|
|
EMP1
|
epithelial membrane protein 1
|
|
chr1_+_181421984
|
7.346
|
|
LAMC2
|
laminin, gamma 2
|
|
chr18_+_19706979
|
6.896
|
NM_000227
NM_001127718
|
LAMA3
|
laminin, alpha 3
|
|
chr11_-_65860450
|
6.435
|
NM_004292
|
RIN1
|
Ras and Rab interactor 1
|
|
chr1_+_181421796
|
5.974
|
NM_005562
NM_018891
|
LAMC2
|
laminin, gamma 2
|
|
chr17_-_36996611
|
5.023
|
NM_000526
|
KRT14
|
keratin 14
|
|
chr1_+_181422016
|
4.612
|
|
LAMC2
|
laminin, gamma 2
|
|
chr3_+_30622905
|
4.569
|
NM_001024847
NM_003242
|
TGFBR2
|
transforming growth factor, beta receptor II (70/80kDa)
|
|
chr1_+_154351082
|
4.375
|
NM_005572
NM_170707
NM_170708
|
LMNA
|
lamin A/C
|
|
chr3_-_48576204
|
4.000
|
NM_033199
|
UCN2
|
urocortin 2
|
|
chr11_+_101488380
|
3.930
|
NM_001195045
|
YAP1
|
Yes-associated protein 1
|
|
chr1_-_151788341
|
3.829
|
NM_002960
|
S100A3
|
S100 calcium binding protein A3
|
|
chr14_-_104507862
|
3.812
|
|
AHNAK2
|
AHNAK nucleoprotein 2
|
|
chr3_-_152517204
|
3.761
|
NM_023915
|
GPR87
|
G protein-coupled receptor 87
|
|
chr9_-_129381086
|
3.723
|
NM_001035534
|
FAM129B
|
family with sequence similarity 129, member B
|
|
chr10_-_88720908
|
3.620
|
|
LOC100133190
|
hypothetical LOC100133190
|
|
chr6_+_83129641
|
3.604
|
NM_006670
|
TPBG
|
trophoblast glycoprotein
|
|
chr1_-_94779727
|
3.559
|
NM_001178096
NM_001993
|
F3
|
coagulation factor III (thromboplastin, tissue factor)
|
|
chr1_-_207891264
|
3.417
|
NM_001017402
|
LAMB3
|
laminin, beta 3
|
|
chr3_-_48607596
|
3.319
|
NM_000094
|
COL7A1
|
collagen, type VII, alpha 1
|
|
chr5_+_140848905
|
3.308
|
NM_018929
NM_032407
|
PCDHGC5
|
protocadherin gamma subfamily C, 5
|
|
chr12_+_76749199
|
3.246
|
NM_014903
|
NAV3
|
neuron navigator 3
|
|
chr3_+_30623376
|
3.184
|
|
TGFBR2
|
transforming growth factor, beta receptor II (70/80kDa)
|
|
chr18_+_59295123
|
3.160
|
NM_002639
|
SERPINB5
|
serpin peptidase inhibitor, clade B (ovalbumin), member 5
|
|
chr1_+_34993234
|
3.113
|
NM_005268
|
GJB5
|
gap junction protein, beta 5, 31.1kDa
|
|
chr12_+_13240983
|
3.080
|
|
EMP1
|
epithelial membrane protein 1
|
|
chr19_-_56215084
|
3.049
|
NM_002776
NM_001077500
|
KLK10
|
kallikrein-related peptidase 10
|
|
chr6_-_152744143
|
3.046
|
|
SYNE1
|
spectrin repeat containing, nuclear envelope 1
|
|
chr6_+_47774247
|
3.020
|
NM_153838
|
GPR115
|
G protein-coupled receptor 115
|
|
chr2_+_28469275
|
3.013
|
NM_005253
|
FOSL2
|
FOS-like antigen 2
|
|
chr7_+_116099648
|
3.009
|
NM_000245
NM_001127500
|
MET
|
met proto-oncogene (hepatocyte growth factor receptor)
|
|
chr16_+_56219802
|
2.996
|
NM_001145771
NM_001145772
NM_001145773
NM_201524
NM_201525
|
GPR56
|
G protein-coupled receptor 56
|
|
chr5_+_150000394
|
2.972
|
NM_001109974
NM_007286
|
SYNPO
|
synaptopodin
|
|
chr18_+_59593591
|
2.969
|
NM_003784
|
SERPINB7
|
serpin peptidase inhibitor, clade B (ovalbumin), member 7
|
|
chr10_+_118177371
|
2.945
|
NM_001011709
|
PNLIPRP3
|
pancreatic lipase-related protein 3
|
|
chr17_-_37022465
|
2.895
|
NM_005557
|
KRT16
|
keratin 16
|
|
chrX_+_99786044
|
2.888
|
|
SRPX2
|
sushi-repeat containing protein, X-linked 2
|
|
chr12_-_56445653
|
2.842
|
|
CYP27B1
|
cytochrome P450, family 27, subfamily B, polypeptide 1
|
|
chr6_-_169392771
|
2.793
|
|
|
|
|
chr15_-_38417564
|
2.735
|
|
C15orf52
|
chromosome 15 open reading frame 52
|
|
chr7_-_42243142
|
2.706
|
NM_000168
|
GLI3
|
GLI family zinc finger 3
|
|
chr4_+_169989680
|
2.641
|
NM_001166110
|
PALLD
|
palladin, cytoskeletal associated protein
|
|
chr13_-_109236897
|
2.633
|
NM_003749
|
IRS2
|
insulin receptor substrate 2
|
|
chr17_-_7962531
|
2.631
|
NM_021628
|
ALOXE3
|
arachidonate lipoxygenase 3
|
|
chr10_+_24795446
|
2.614
|
|
KIAA1217
|
KIAA1217
|
|
chr3_+_11153778
|
2.603
|
NM_001098213
|
HRH1
|
histamine receptor H1
|
|
chrX_+_149364377
|
2.592
|
NM_001177466
NM_005491
|
MAMLD1
|
mastermind-like domain containing 1
|
|
chr4_-_115120251
|
2.581
|
NM_024590
|
ARSJ
|
arylsulfatase family, member J
|
|
chr12_-_51173238
|
2.575
|
NM_005554
|
KRT6A
|
keratin 6A
|
|
chr19_+_43572679
|
2.558
|
NM_001039616
NM_001042522
|
SPRED3
|
sprouty-related, EVH1 domain containing 3
|
|
chrX_+_99785818
|
2.525
|
NM_014467
|
SRPX2
|
sushi-repeat containing protein, X-linked 2
|
|
chr5_-_141041982
|
2.515
|
NM_022481
|
ARAP3
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 3
|
|
chr17_-_24068759
|
2.487
|
NM_001144942
NM_031934
|
RAB34
|
RAB34, member RAS oncogene family
|
|
chr13_-_37070862
|
2.431
|
NM_001135934
NM_001135935
NM_001135936
NM_006475
|
POSTN
|
periostin, osteoblast specific factor
|
|
chr20_-_1257782
|
2.429
|
|
SDCBP2
|
syndecan binding protein (syntenin) 2
|
|
chr12_-_105057940
|
2.419
|
NM_014840
|
NUAK1
|
NUAK family, SNF1-like kinase, 1
|
|
chr5_+_145297490
|
2.414
|
|
SH3RF2
|
SH3 domain containing ring finger 2
|
|
chr5_-_141041933
|
2.409
|
|
ARAP3
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 3
|
|
chr6_-_152744022
|
2.404
|
|
SYNE1
|
spectrin repeat containing, nuclear envelope 1
|
|
chr5_+_176807931
|
2.349
|
NM_001174102
|
PRR7
|
proline rich 7 (synaptic)
|
|
chr16_-_2848106
|
2.341
|
NM_022119
|
PRSS22
|
protease, serine, 22
|
|
chr20_+_19903778
|
2.341
|
|
RIN2
|
Ras and Rab interactor 2
|
|
chr9_+_123369219
|
2.334
|
NM_032552
|
DAB2IP
|
DAB2 interacting protein
|
|
chr3_-_150578207
|
2.331
|
NM_014220
|
TM4SF1
|
transmembrane 4 L six family member 1
|
|
chr7_-_42243318
|
2.298
|
|
GLI3
|
GLI family zinc finger 3
|
|
chr3_+_57069508
|
2.270
|
NM_181727
|
SPATA12
|
spermatogenesis associated 12
|
|
chr17_-_24069299
|
2.262
|
NM_001142624
NM_001142625
NM_001144943
|
RAB34
|
RAB34, member RAS oncogene family
|
|
chr3_-_150578103
|
2.261
|
|
TM4SF1
|
transmembrane 4 L six family member 1
|
|
chr1_+_154351162
|
2.258
|
|
LMNA
|
lamin A/C
|
|
chr3_-_150578148
|
2.218
|
|
TM4SF1
|
transmembrane 4 L six family member 1
|
|
chr17_+_71229151
|
2.180
|
|
ITGB4
|
integrin, beta 4
|
|
chr17_-_37034288
|
2.179
|
NM_000422
|
KRT17
|
keratin 17
|
|
chr2_+_28469199
|
2.162
|
|
FOSL2
|
FOS-like antigen 2
|
|
chr5_+_145296313
|
2.148
|
NM_152550
|
SH3RF2
|
SH3 domain containing ring finger 2
|
|
chr16_-_46823682
|
2.145
|
NM_033151
|
ABCC11
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 11
|
|
chr5_+_35892747
|
2.143
|
NM_002185
|
IL7R
|
interleukin 7 receptor
|
|
chr11_+_12265022
|
2.126
|
NM_032867
|
MICALCL
|
MICAL C-terminal like
|
|
chr4_+_81325447
|
2.125
|
NM_020226
|
PRDM8
|
PR domain containing 8
|
|
chr17_+_71229110
|
2.119
|
NM_000213
NM_001005731
|
ITGB4
|
integrin, beta 4
|
|
chr18_+_59705918
|
2.113
|
NM_001143818
NM_002575
|
SERPINB2
|
serpin peptidase inhibitor, clade B (ovalbumin), member 2
|
|
chr11_-_128399218
|
2.080
|
NM_014715
|
ARHGAP32
|
Rho GTPase activating protein 32
|
|
chr19_+_7859380
|
2.075
|
NM_025061
|
LRRC8E
|
leucine rich repeat containing 8 family, member E
|
|
chr11_-_117640196
|
2.073
|
|
MPZL2
|
myelin protein zero-like 2
|
|
chr17_+_21131940
|
2.072
|
NM_002756
|
MAP2K3
|
mitogen-activated protein kinase kinase 3
|
|
chr11_-_66898223
|
2.049
|
NM_001166212
|
CLCF1
|
cardiotrophin-like cytokine factor 1
|
|
chr10_+_24795465
|
2.048
|
|
KIAA1217
|
KIAA1217
|
|
chr9_+_5500544
|
2.041
|
NM_025239
|
PDCD1LG2
|
programmed cell death 1 ligand 2
|
|
chr7_+_48094750
|
2.038
|
NM_181597
|
UPP1
|
uridine phosphorylase 1
|
|
chr22_-_28972683
|
2.036
|
NM_002309
|
LIF
|
leukemia inhibitory factor (cholinergic differentiation factor)
|
|
chr3_-_188936941
|
2.035
|
NM_001130845
|
BCL6
|
B-cell CLL/lymphoma 6
|
|
chr11_+_35154775
|
1.991
|
|
CD44
|
CD44 molecule (Indian blood group)
|
|
chr14_-_22515725
|
1.988
|
|
JUB
|
jub, ajuba homolog (Xenopus laevis)
|
|
chr15_+_38319326
|
1.985
|
NM_020168
|
PAK6
|
p21 protein (Cdc42/Rac)-activated kinase 6
|
|
chr11_-_6298284
|
1.963
|
NM_145040
|
PRKCDBP
|
protein kinase C, delta binding protein
|
|
chr21_-_35343331
|
1.956
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr17_+_71229191
|
1.947
|
|
ITGB4
|
integrin, beta 4
|
|
chr15_-_68781662
|
1.923
|
NM_001008224
|
UACA
|
uveal autoantigen with coiled-coil domains and ankyrin repeats
|
|
chr11_+_696607
|
1.919
|
|
EPS8L2
|
EPS8-like 2
|
|
chr6_+_31190505
|
1.910
|
NM_014068
|
PSORS1C1
|
psoriasis susceptibility 1 candidate 1
|
|
chr17_-_36928604
|
1.894
|
NM_002275
|
KRT15
|
keratin 15
|
|
chr2_+_28469172
|
1.894
|
|
FOSL2
|
FOS-like antigen 2
|
|
chr6_+_45404031
|
1.892
|
NM_001015051
NM_001024630
|
RUNX2
|
runt-related transcription factor 2
|
|
chr14_+_51025604
|
1.861
|
NM_001042481
|
FRMD6
|
FERM domain containing 6
|
|
chr11_-_119499188
|
1.858
|
|
TRIM29
|
tripartite motif containing 29
|
|
chr13_+_77007809
|
1.851
|
NM_001160706
NM_003843
NM_144777
|
SCEL
|
sciellin
|
|
chr21_-_35343456
|
1.842
|
NM_001754
|
RUNX1
|
runt-related transcription factor 1
|
|
chr5_-_1935764
|
1.836
|
NM_016358
|
IRX4
|
iroquois homeobox 4
|
|
chr14_+_95792299
|
1.836
|
NM_000710
|
BDKRB1
|
bradykinin receptor B1
|
|
chr2_-_85494664
|
1.821
|
|
CAPG
|
capping protein (actin filament), gelsolin-like
|
|
chr2_+_233605625
|
1.812
|
NM_005383
|
NEU2
|
sialidase 2 (cytosolic sialidase)
|
|
chr11_+_384199
|
1.795
|
NM_007183
|
PKP3
|
plakophilin 3
|
|
chr7_-_1465487
|
1.793
|
|
MICALL2
|
MICAL-like 2
|
|
chr1_+_154351200
|
1.791
|
|
LMNA
|
lamin A/C
|
|
chr6_+_12398514
|
1.777
|
NM_001168319
NM_001955
|
EDN1
|
endothelin 1
|
|
chr2_-_73170290
|
1.772
|
|
RAB11FIP5
|
RAB11 family interacting protein 5 (class I)
|
|
chr8_+_32623792
|
1.769
|
NM_013959
|
NRG1
|
neuregulin 1
|
|
chr11_+_384238
|
1.769
|
|
PKP3
|
plakophilin 3
|
|
chr9_-_116920232
|
1.766
|
NM_002160
|
TNC
|
tenascin C
|
|
chr1_-_115682374
|
1.754
|
NM_002506
|
NGF
|
nerve growth factor (beta polypeptide)
|
|
chr21_-_35343448
|
1.740
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr9_+_131974506
|
1.729
|
NM_014286
|
NCS1
|
neuronal calcium sensor 1
|
|
chr1_+_151147646
|
1.722
|
NM_005547
|
IVL
|
involucrin
|
|
chr1_-_150564302
|
1.697
|
NM_002016
|
FLG
|
filaggrin
|
|
chr11_-_102174102
|
1.696
|
NM_001145938
NM_002421
|
MMP1
|
matrix metallopeptidase 1 (interstitial collagenase)
|
|
chr21_-_35181956
|
1.677
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr19_-_18912112
|
1.677
|
NM_001145722
NM_001145721
|
HOMER3
|
homer homolog 3 (Drosophila)
|
|
chr6_+_121798443
|
1.653
|
NM_000165
|
GJA1
|
gap junction protein, alpha 1, 43kDa
|
|
chr3_+_189413414
|
1.642
|
NM_005578
|
LPP
|
LIM domain containing preferred translocation partner in lipoma
|
|
chr11_-_102174032
|
1.642
|
|
MMP1
|
matrix metallopeptidase 1 (interstitial collagenase)
|
|
chr8_+_120289735
|
1.636
|
NM_052886
|
MAL2
|
mal, T-cell differentiation protein 2 (gene/pseudogene)
|
|
chr1_-_177378801
|
1.615
|
NM_001136000
NM_001168239
NM_005158
|
ABL2
|
v-abl Abelson murine leukemia viral oncogene homolog 2
|
|
chr5_+_73145098
|
1.613
|
|
RGNEF
|
190 kDa guanine nucleotide exchange factor
|
|
chr1_-_151280201
|
1.609
|
NM_006945
|
SPRR2D
|
small proline-rich protein 2D
|
|
chr21_+_46226072
|
1.604
|
NM_001848
|
COL6A1
|
collagen, type VI, alpha 1
|
|
chr19_-_43955984
|
1.604
|
NM_002307
|
LGALS7
LGALS7B
|
lectin, galactoside-binding, soluble, 7
lectin, galactoside-binding, soluble, 7B
|
|
chr11_+_696103
|
1.599
|
NM_022772
|
EPS8L2
|
EPS8-like 2
|
|
chr12_-_46439127
|
1.598
|
NM_001098531
|
RAPGEF3
|
Rap guanine nucleotide exchange factor (GEF) 3
|
|
chr3_-_31997738
|
1.594
|
|
OSBPL10
|
oxysterol binding protein-like 10
|
|
chr3_-_31998241
|
1.593
|
NM_001174060
NM_017784
|
OSBPL10
|
oxysterol binding protein-like 10
|
|
chr2_-_166359016
|
1.590
|
NM_004482
|
GALNT3
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 3 (GalNAc-T3)
|
|
chrX_-_10761729
|
1.571
|
NM_033290
|
MID1
|
midline 1 (Opitz/BBB syndrome)
|
|
chr10_+_134751398
|
1.561
|
NM_001083909
|
GPR123
|
G protein-coupled receptor 123
|
|
chr20_+_33223434
|
1.533
|
NM_006404
|
PROCR
|
protein C receptor, endothelial
|
|
chr2_+_113591940
|
1.531
|
NM_000577
NM_173841
NM_173843
|
IL1RN
|
interleukin 1 receptor antagonist
|
|
chr8_+_22900863
|
1.530
|
NM_001160036
|
RHOBTB2
|
Rho-related BTB domain containing 2
|
|
chr2_-_85494639
|
1.519
|
|
CAPG
|
capping protein (actin filament), gelsolin-like
|
|
chr11_+_47386621
|
1.519
|
NM_001128225
NM_152264
|
SLC39A13
|
solute carrier family 39 (zinc transporter), member 13
|
|
chr11_-_8695974
|
1.507
|
|
ST5
|
suppression of tumorigenicity 5
|
|
chr11_+_35596310
|
1.503
|
NM_014344
|
FJX1
|
four jointed box 1 (Drosophila)
|
|
chr9_-_109291866
|
1.480
|
NM_004235
|
KLF4
|
Kruppel-like factor 4 (gut)
|
|
chr6_+_41712907
|
1.479
|
|
MDFI
|
MyoD family inhibitor
|
|
chr2_-_18634257
|
1.469
|
NM_001002006
NM_033253
|
NT5C1B-RDH14
NT5C1B
|
NT5C1B-RDH14 readthrough
5'-nucleotidase, cytosolic IB
|
|
chr11_+_20005704
|
1.458
|
|
NAV2
|
neuron navigator 2
|
|
chr6_-_116799548
|
1.454
|
|
LOC100287467
|
hypothetical LOC100287467
|
|
chr20_+_44070946
|
1.450
|
NM_004994
|
MMP9
|
matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase)
|
|
chr1_+_15128882
|
1.449
|
NM_001018001
|
KAZN
|
kazrin, periplakin interacting protein
|
|
chr19_-_48661563
|
1.448
|
NM_014400
|
LYPD3
|
LY6/PLAUR domain containing 3
|
|
chr2_+_220200530
|
1.439
|
NM_005070
NM_201574
|
SLC4A3
|
solute carrier family 4, anion exchanger, member 3
|
|
chr15_+_84021271
|
1.419
|
NM_144767
|
AKAP13
|
A kinase (PRKA) anchor protein 13
|
|
chr6_-_138470160
|
1.418
|
NM_022121
|
PERP
|
PERP, TP53 apoptosis effector
|
|
chr18_+_50646616
|
1.413
|
NM_004163
|
RAB27B
|
RAB27B, member RAS oncogene family
|
|
chr16_+_55199978
|
1.410
|
NM_005953
|
MT2A
|
metallothionein 2A
|
|
chr21_-_35343065
|
1.409
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr18_+_59788242
|
1.394
|
NM_001031848
NM_002640
NM_198833
|
SERPINB8
|
serpin peptidase inhibitor, clade B (ovalbumin), member 8
|
|
chr10_+_102782151
|
1.392
|
|
SFXN3
|
sideroflexin 3
|
|
chr9_-_34652629
|
1.389
|
NM_006664
|
CCL27
|
chemokine (C-C motif) ligand 27
|
|
chr21_-_35343500
|
1.379
|
|
RUNX1
|
runt-related transcription factor 1
|
|
chr19_+_49866563
|
1.357
|
NM_001127893
NM_020219
|
CEACAM19
|
carcinoembryonic antigen-related cell adhesion molecule 19
|
|
chr5_+_36642424
|
1.353
|
|
SLC1A3
|
solute carrier family 1 (glial high affinity glutamate transporter), member 3
|
|
chr12_-_46438447
|
1.348
|
NM_001098532
NM_006105
|
RAPGEF3
|
Rap guanine nucleotide exchange factor (GEF) 3
|
|
chrX_+_106030049
|
1.336
|
NM_001171092
|
CLDN2
|
claudin 2
|
|
chr1_-_33139425
|
1.325
|
NM_033504
|
TMEM54
|
transmembrane protein 54
|
|
chr15_+_38318528
|
1.319
|
|
PAK6
|
p21 protein (Cdc42/Rac)-activated kinase 6
|
|
chr16_+_88515917
|
1.317
|
NM_001197181
|
TUBB3
|
tubulin, beta 3
|
|
chr11_-_123353607
|
1.315
|
NM_001004474
|
OR10S1
|
olfactory receptor, family 10, subfamily S, member 1
|
|
chr11_+_55335418
|
1.306
|
NM_001004738
|
OR5L1
|
olfactory receptor, family 5, subfamily L, member 1
|
|
chr5_+_52812236
|
1.301
|
NM_006350
NM_013409
|
FST
|
follistatin
|
|
chr1_-_84237316
|
1.298
|
NM_024686
|
TTLL7
|
tubulin tyrosine ligase-like family, member 7
|
|
chr7_+_28692122
|
1.297
|
NM_001011666
|
CREB5
|
cAMP responsive element binding protein 5
|
|
chr1_+_27062215
|
1.294
|
NM_006142
|
SFN
|
stratifin
|
|
chr11_-_47164001
|
1.293
|
NM_001184975
|
PACSIN3
|
protein kinase C and casein kinase substrate in neurons 3
|
|
chr2_-_142605010
|
1.287
|
|
LRP1B
|
low density lipoprotein receptor-related protein 1B
|
|
chr6_+_74462654
|
1.280
|
|
CD109
|
CD109 molecule
|
|
chr10_-_30064600
|
1.263
|
NM_003174
|
SVIL
|
supervillin
|
|
chr10_-_118491971
|
1.258
|
NM_025015
|
HSPA12A
|
heat shock 70kDa protein 12A
|
|
chr1_+_17507276
|
1.251
|
NM_012387
|
PADI4
|
peptidyl arginine deiminase, type IV
|
|
chr11_-_117640166
|
1.249
|
|
MPZL2
|
myelin protein zero-like 2
|
|
chr11_+_47386707
|
1.245
|
|
SLC39A13
|
solute carrier family 39 (zinc transporter), member 13
|
|
chr10_-_93382837
|
1.240
|
NM_005398
|
PPP1R3C
|
protein phosphatase 1, regulatory (inhibitor) subunit 3C
|
|
chr1_+_35019376
|
1.239
|
NM_024009
|
GJB3
|
gap junction protein, beta 3, 31kDa
|
|
chr18_+_54039818
|
1.221
|
|
NEDD4L
|
neural precursor cell expressed, developmentally down-regulated 4-like
|
|
chr2_-_65447267
|
1.219
|
|
SPRED2
|
sprouty-related, EVH1 domain containing 2
|
|
chr15_-_53996597
|
1.213
|
NM_198400
|
NEDD4
|
neural precursor cell expressed, developmentally down-regulated 4
|
|
chr1_-_153272859
|
1.211
|
NM_144622
|
DCST2
|
DC-STAMP domain containing 2
|
|
chr21_-_26465005
|
1.206
|
NM_000484
NM_001136129
NM_001136130
NM_201413
NM_201414
|
APP
|
amyloid beta (A4) precursor protein
|
|
chr21_-_26464931
|
1.203
|
|
APP
|
amyloid beta (A4) precursor protein
|
|
chr4_+_108965167
|
1.199
|
NM_001136258
|
SGMS2
|
sphingomyelin synthase 2
|
|
chr12_-_54646061
|
1.196
|
NM_006928
|
PMEL
|
premelanosome protein
|
|
chr17_-_3408038
|
1.196
|
NM_145068
|
TRPV3
|
transient receptor potential cation channel, subfamily V, member 3
|